Created computer based on DNA, which finally can be reprogrammed
 Source:
Source:
It Is believed that DNA will save us from computers. Thanks to advances in the replacement of silicon transistors, computers based on DNA promise to provide us a massive parallel computing architecture, is impossible at the present time. But here's the catch: the molecular circuits that were created before today, had absolutely no flexibility. Today, the use of DNA for computing is the same as the "create new computer from new hardware to run only one program," says David Doty.
Doty, Professor, University of California at Davis, and his colleagues decided to find out what it would take to create a DNA computer that actually can be reprogrammed.
theComputer from DNA
In an article published this week in the journal Nature, Doty and his colleagues at the University of California and the University of Maynooth has demonstrated just that. They showed that it is possible to use a simple trigger to make the same basic set of DNA molecules to implement many different algorithms. Although this study is still exploratory in nature, in the future can be used reprogrammable algorithms for molecular programming DNA robots that have successfully delivered drugs to the cancer cells.
"This is one of the most important works in the field," says Thorsten-Lars Schmidt, associate Professor of experimental Biophysics at Kent state University who did not participate in the study. "Earlier it was algorithmic self-Assembly, but not to this degree of complexity".
In the electronic computers like the fact that you are using to read this article, bits — binary units of information that tell the computer what to do. They represent discrete physical state of the underlying hardware, usually in the form of the presence or absence of electric current. These bits — or even an electric signal which is transmitted through the circuit consisting of logic elements that perform an operation on one or more input bits and produce one bit as output.
Combining these simple building blocks again and again, computers can run a surprisingly complex program. The idea behind DNA-based computing, is to replace the chemical bonds electrical signals and nucleic acid — silicon, and to create biomolecular software. According to Eric Winfrey, a computer scientist from Caltech and co-author of the work, molecular algorithms use the natural ability of information processing, sewn into the DNA, but instead to give control to nature, "the growth process is controlled by computers".
Over the past 20 years in several experiments, we used molecular algorithms for things such as TIC-TAC-toe or Assembly of various shapes. In each of these cases, the DNA sequence had to be carefully designed to create one specific algorithm that would generate the structure of DNA. What's different in this case is the fact that researchers have developed a system in which the same underlying DNA fragments can be sequenced to create an entirely different algorithms and therefore can be made of different end products.
This process begins with DNA origami, a method of folding a long piece of DNA into a desired shape. This rolled-up piece of DNA serves as a "led" (seed, seed), which runs an algorithmic pipeline, like on a string dipped in sugared water, gradually grows the caramel. The seed remains largely the same irrespective of the algorithm and changes are made only in a few small sequences for each new experiment.
Once the scientists have created the seed, they added in a solution of 100 other strands of DNA, DNA fragments. These fragments, each of which consists of a unique arrangement 42 of the nucleic bases (the four basic biological molecules that comprise DNA) taken from a large collection of 355 fragments of DNA created by scientists. To create another algorithm scientists have to choose a different set of starting fragments. Molecular algorithm that includes random walk requires a different set of DNA fragments that the algorithm uses for calculation. Since these DNA fragments are joined in the Assembly process, they form a circuit that implements the selected molecular algorithm on the input bits provided by the led.
Using this system, scientists have created 21 different algorithm, which can perform tasks such as recognizing a multiple of three, the choice of leader, generation patterns, and the score to 63. All these algorithms were implemented using different combinations of the same 355 DNA fragments.
Of Course, to write code by resetting of DNA fragments in a test tube while it does not, however, the whole idea is a model for future iterations of flexible computers based on DNA. If Doty, Winfrey and woods get their way, molecular programmers of tomorrow even won't think about the biomechanics underlying their programs, just as modern programmers do not have to understand the physics of transistors to write good software.
The Potential use of this nano-scale Assembly technique of the imagination, but these projections are based on our relatively limited understanding of the nanoscaleworld. Alan Turing was not able to predict the advent of the Internet, and therefore we may have incomprehensible the application of molecular Informatics.
What will be capable of molecular computers? Tell us .
Recommended
Is it possible digital immortality and whether it
when will man become immortal through digital technologies. I don't believe it. And you? In 2016, the youngest daughter Jang JI-sen This died of the disease associated with the blood. But in February, the mother was reunited with her daughter in virt...
Why bad long sit at the computer and how to fix it
I've recently conducted a small survey among friends and acquaintances about how they evaluate their effectiveness when working remotely. Almost everyone I know — now work from home with computer and phone. And, as it turned out, even those who...
Parametric architecture: can artificial intelligence to design cities?
When you think about the future, what pictures arise in front of your eyes? As a lover of retro-futurism – a genre which is based on representation of the people in the past about the future, I always imagined the city of the future built buildings, ...
Related News
Quantum computers. Why them yet, although they already have?
Fifty years ago, smartphones would have seemed absolutely magical computers. Just as classical computers have been almost unimaginable to previous generations, today we are facing the birth of an entirely new type of computing: so...
IBM invented "Moore's law" for quantum computers
IBM has proposed the use of a measure of the "quantum volume", which is expected to double every year — and it will be the equivalent of Moore's law, which is observed in traditional computing. According to Moore's law the number ...
The fastest supercomputer in the world broke the record of artificial intelligence
On the West coast of America the most valuable company in the world trying to make artificial intelligence smarter. Google and Facebook brag experiments using billions of photos and thousands of high-performance processors. But at...
Physicists have calculated the time of the state of superposition of graphene qubits
the Possibility of practical use of quantum computers one step closer thanks to graphene. Experts from the Massachusetts Institute of technology and their colleagues from other research institutions were able to calculate the time...
At MIT used a biological virus in order to speed up your computer
whenever the computer (and any other electronic device) processes the data, there is a small delay, that is to say, the transfer of information "from one equipment to another" (e.g. from the memory to physical). The more powerful ...
New particles could open the way to photonic computers
All modern electronic devices use to transmit information of the electrons. Now in full swing, the development of quantum computers, which many consider the future replacement of traditional devices. However, there is another, no ...
New computer type architecture of the brain can improve data processing methods
Scientists from IBM are developing a new computer architecture that will be better suited to handle increasing volumes of data coming from algorithms of artificial intelligence. They draw their inspiration from the human brain, an...
The company D-Wave has launched an open and free platform for quantum computing
With the wide spread of quantum computers needs to produce a real revolution in the field of computer science, providing not only extra power but also improved performance in cybersecurity. We already have quantum computers, but ...
Excursion to the Museum of computers that changed the world
For some reason, old computers don't become classics. Few people contains them with the same concern as contain antique furniture or cars. Probably the reason that they are not suitable for use in the modern world even though the ...
The Intel found 3 vulnerabilities. They allow you to steal
Today Intel announced three new vulnerabilities of their processors. According to the American company, these vulnerabilities can be exploited to gain access to some data stored in the computer memory. Under the threat of a proces...
9-th generation Intel CPU with 8 cores will be presented October 1
there were rumors that Intel introduced the 9 generation of processors in October. Although the 10-nanometer chips Cannon Lake company was postponed until 2019, the updates this year will be based on the improvement of the existin...
The story of the first Macintosh computer, which is the headquarters of Microsoft
Not so long ago the history of the first business card of bill gates and Paul Allen, which is stored in the exhibition hall of the headquarters of Microsoft are available to visit. In this room there's something else worthy of not...
The story of the first business card of bill gates and Paul Allen
All with something started. Microsoft started with two friends who decided to write software for the microcomputer. They started a company and made myself business cards. Today, these business cards are stored in the Microsoft hea...
Users of computers began to threaten the publication of personal videos and browser history
computer Users around the world have started to receive e-mails from Scam to extort money. Depending on who sent the letter, its contents may change. However, colleagues from Business Insider was able to extract common features in...
7 laptops for those who want to buy the best
If you want to buy the most efficient laptop today, you need to look at the model with a premium design and materials, high-resolution display, processor and 16 gigabytes of RAM. In this case, you have the choice not too big. Here...
The hacker stole the files of the U.S. army, but could not sell them even for $ 150
the Hacker has guessed to use the vulnerability of routers to gain access to the files of the U.S. army. The obtained data he attempted to sell on the forum on the darknet, but was unable to find an interested buyer, even reducing...
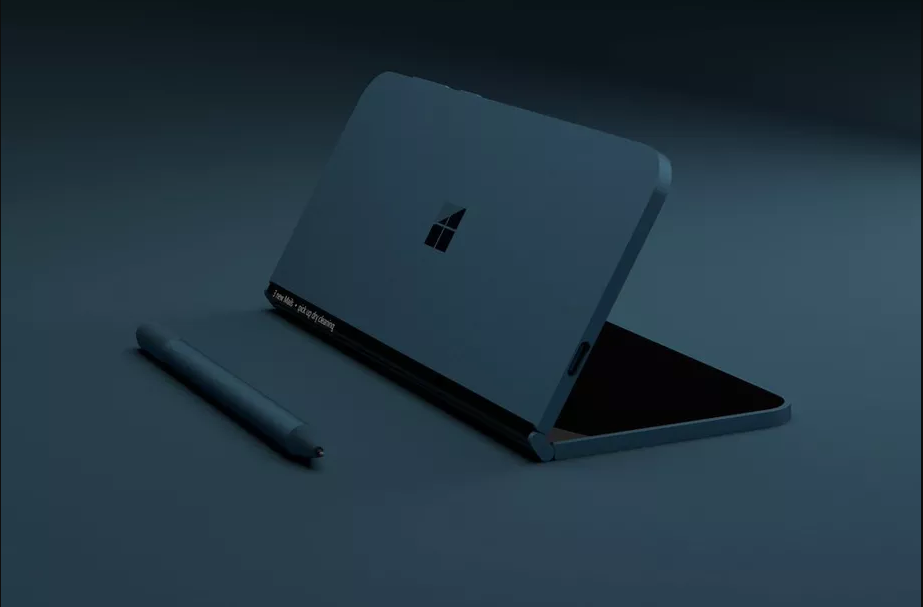
Secret "pocket" Surface from Microsoft is with a folding screen
Microsoft is working on a mysterious new device from the Surface has at least a few years. The device, codenamed "Andromeda" was repeatedly flashed in patents, reports, references in the operating system, and should include design...
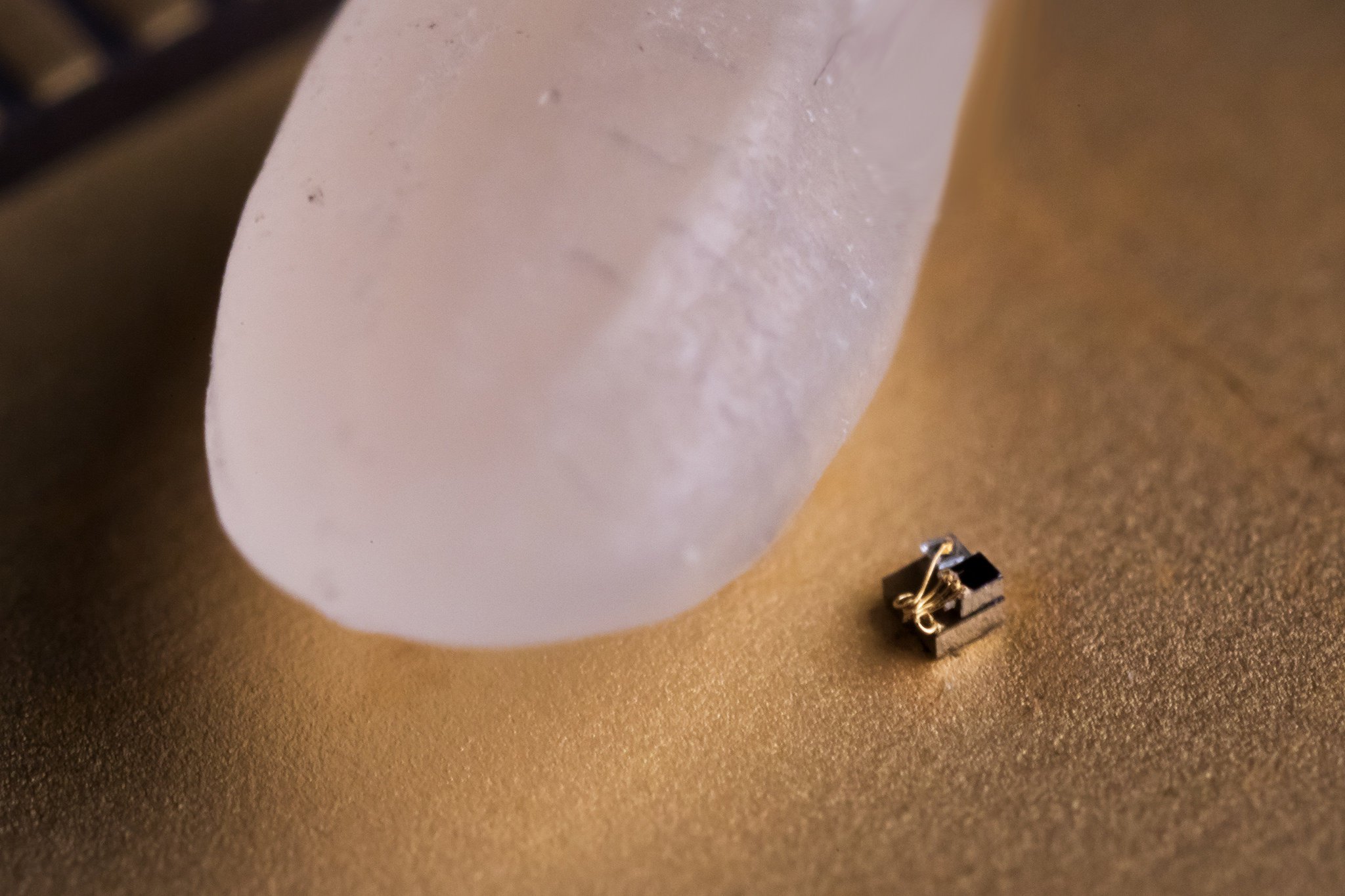
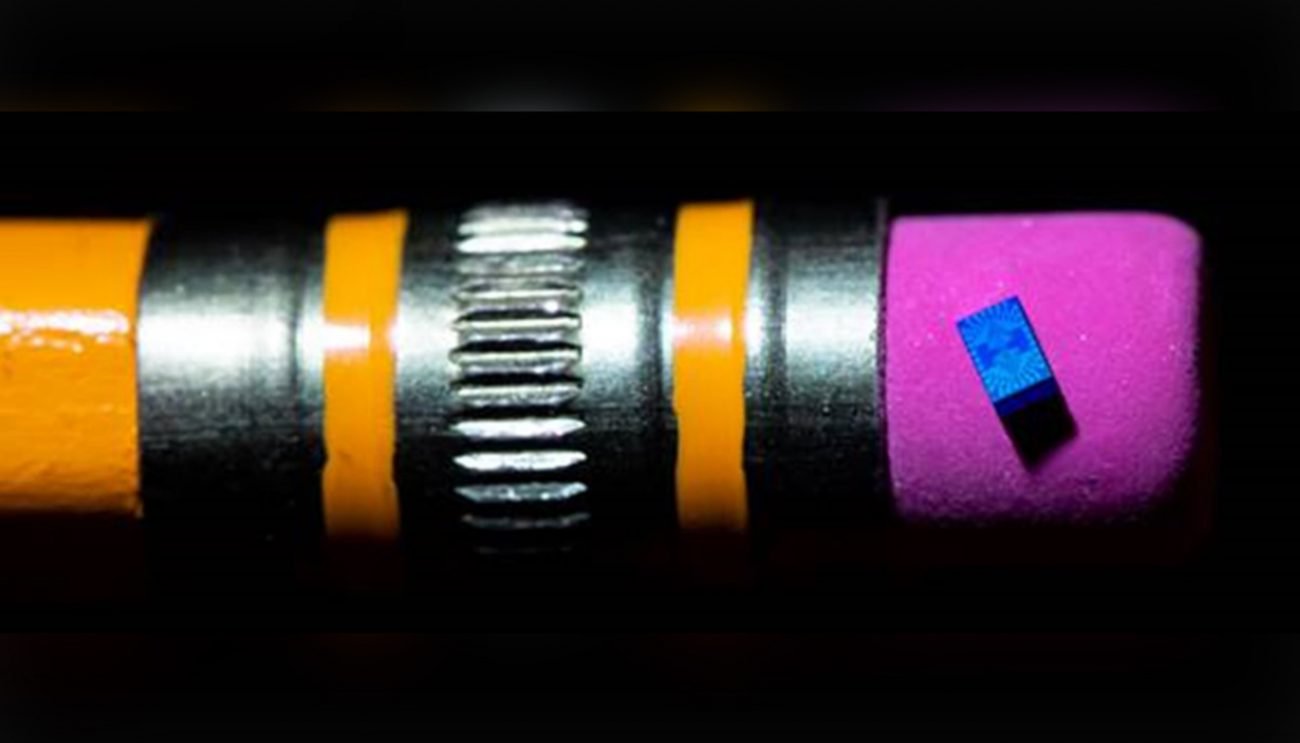
The smallest computer in the world has become even smaller
When IBM said in March that created the smallest computer in the world, scientists from the University of Michigan with displeasure raised eyebrows because IBM is the championship belonged to them. Now the smallest computer in the...
Intel has created the world's smallest quantum chip
Already practically does not remain doubts that in the future quantum computers, if not will replace the familiar to us computers, then certainly strongly they will press. The success in this sphere do have a company and recently,...
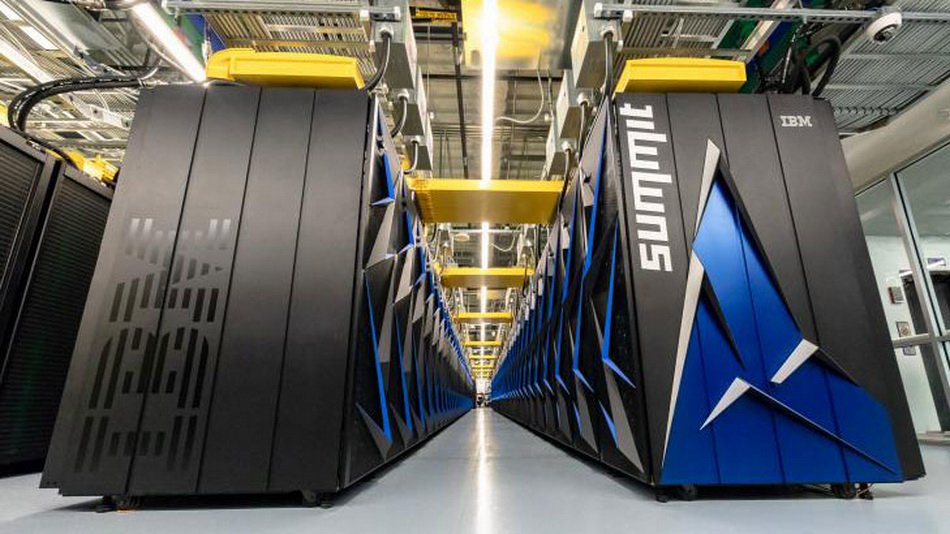
In the United States represented the most powerful supercomputer in the world
the US Department of energy introduced the world's most powerful supercomputer, selected the title from the Chinese Sunway TaihuLight. Peak performance supercomputer American Summit reached 200 petaflops. (200 quadrillion calculat...








































Comments (0)
This article has no comment, be the first!